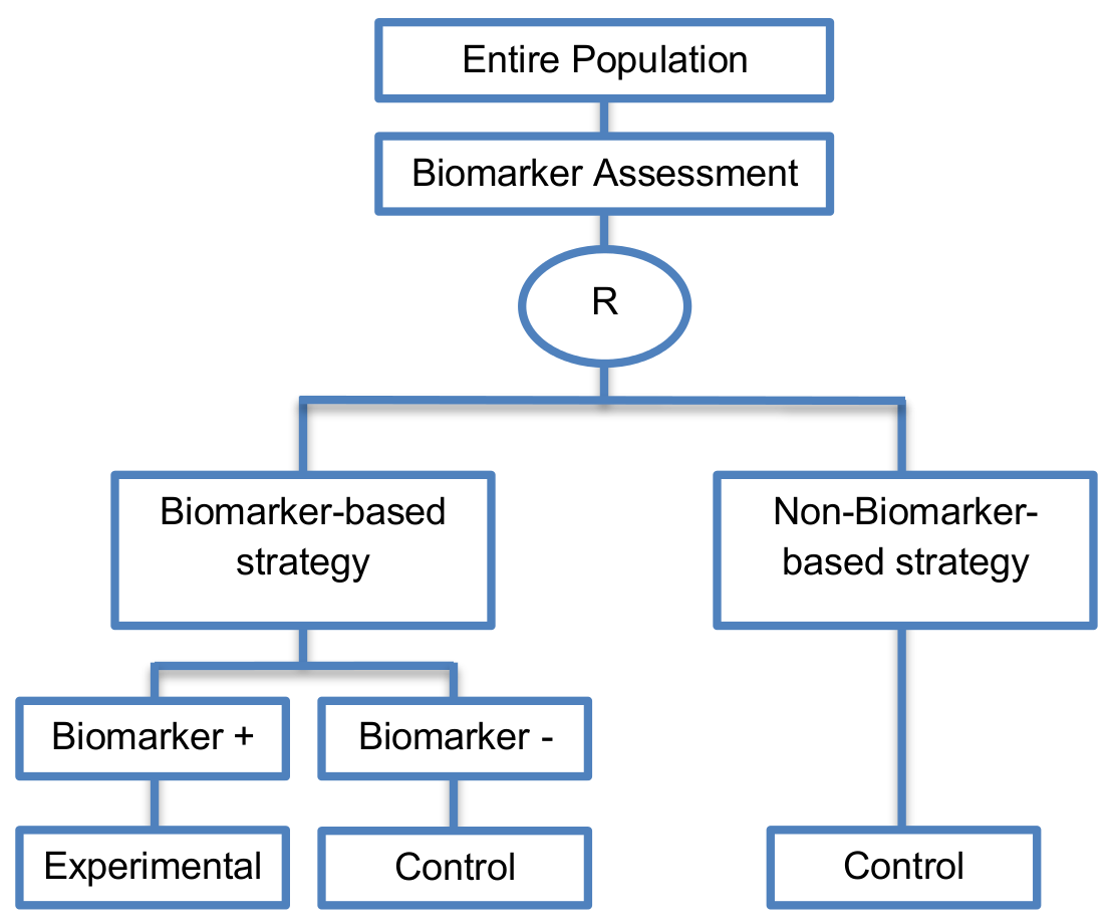
In the biomarker-based strategy arm, biomarker-positive patients receive the experimental treatment, whereas, biomarker-negative patients receive the control treatment. Patients who are randomized to the non-biomarker-based strategy arm receive the control treatment irrespective of their biomarker status.
Alternative names: Marker strategy designs, Biomarker-strategy designs, Strategy design, Marker-based strategy designs, Marker-based designs, Random disclosure designs, Customized strategy designs, Parallel controlled pharmacogenetic study designs, Marker-based strategy designs I, Biomarker-guided designs, Biomarker-based assignment of specific drug therapy designs, Marker-based strategy I designs, Biomarker-strategy designs with a standard control, Marker strategy designs for prognostic biomarkers
Details
Utility
This approach is useful when we want to test the hypothesis that the treatment effect based on the biomarker-based strategy approach is superior to that of the standard of care.
Methodology
- The clinical utility of a biomarker can be evaluated by comparing the two strategy groups.
- The predictive utility of the marker-based treatment strategy could be assessed by comparing the outcome of all patients in the biomarker-based strategy arm to all patients in the non-biomarker-based strategy arm.
- Patients in the marker-based strategy arm do not need to be limited to two treatments; in principle, a marker-based strategy involving many biomarkers and many possible treatments could be compared to standard of care treatment.
Sample size Formula
-
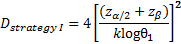
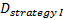 is referred to the required
total number of events (time-to-event outcome),
is referred to the required
total number of events (time-to-event outcome),
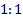 ratio between the two treatment
arms (experimental:control) is assumed.
ratio between the two treatment
arms (experimental:control) is assumed.
-
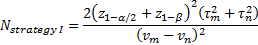
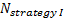 is referred to the required
total sample size (continuous clinical endpoints),
is referred to the required
total sample size (continuous clinical endpoints),
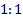 ratio between the two treatment
arms (experimental:control) is assumed,
ratio between the two treatment
arms (experimental:control) is assumed,
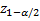 ,
,
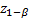 denote the lower
denote the lower
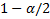 - and lower
- and lower
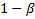 -points respectively of a
standard normal distribution,
-points respectively of a
standard normal distribution,
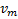 and
and
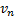 denote the mean response from
the biomarker-based strategy arm and the non-biomarker-based strategy arm
respectively, and
denote the mean response from
the biomarker-based strategy arm and the non-biomarker-based strategy arm
respectively, and
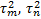 denote the variance of response
for the biomarker-based strategy arm and non-biomarker-based strategy arm
respectively.
denote the variance of response
for the biomarker-based strategy arm and non-biomarker-based strategy arm
respectively.
-
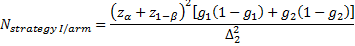
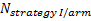 is referred to the required
total number of patients per arm (binary outcome),
is referred to the required
total number of patients per arm (binary outcome),
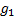 is the expected response rate in
the biomarker-based strategy arm,
is the expected response rate in
the biomarker-based strategy arm,
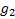 is the expected response rate in
the non biomarker-based strategy arm,
is the expected response rate in
the non biomarker-based strategy arm,
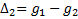 ,
,
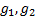 can be found by calculating the
formulae
can be found by calculating the
formulae
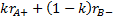 and
and
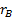 respectively,
respectively,
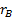 denotes the marginal effect of
treatment B (control treatment).
denotes the marginal effect of
treatment B (control treatment).
Statistical/Practical considerations
Advantages
- Biomarker can be validated without including all possible biomarker–treatment combinations (in the non-biomarker-based arm all patients receive only the control treatment instead of receiving both the experimental and control treatment).
- Have the option of testing the biomarker status of patients in the non-biomarker-strategy arm which can aid secondary analyses.
- Able to inform us whether the biomarker is prognostic.
- Can be expanded to investigate several biomarkers and treatments. Additionally, these designs can be attractive when evaluating multiple biomarkers or the predictive value of molecular profiling between several treatment options is to be assessed.
- Might be used more frequently in the future due to the wide variety of molecular biomarkers, complexity of gene expression arrays, and several treatments directed at similar targets.
Limitations
- Unable to inform us whether the biomarker is predictive as these designs are able to answer the question about whether the biomarker-based strategy is more effective than standard treatment, irrespective of the biomarker status of the study population.
- The evaluation of the true biomarker by treatment effect is not possible as the biomarker-positive patients receive only the experimental treatment and not the alternative treatment (control treatment). Consequently, these designs cannot detect the case in which the control treatment might be more beneficial for the entire population.
- In case that the number of biomarker-positive patients is very small, then the treatment received will be similar in biomarker-strategy arm and non-biomarker strategy arm. Consequently, the trial might give little information regarding the efficacy of the experimental treatment or it might not be able to detect it. As a result, these types of designs should be used when there is an adequate number of biomarker-positive and biomarker-negative patients.
- Unable to compare directly experimental treatment to control treatment as the aim is to compare not the treatments but the biomarker-strategies.
- Less efficient designs than biomarker-stratified designs and a poor substitute for clinical trials which aim to compare the experimental treatment to control treatment, since it is possible for some patients in both the biomarker-based strategy arm and non-biomarker-based strategy arm to be assigned to the same treatment (due to the existence of biomarker-negative patients in both strategy arms the treatment effect can be diluted). Consequently, as a large overlap of patients receiving the same treatment might have occurred, the comparison of the two biomarker-strategy arms results in a hazard ratio which is forced towards unity, i.e., no treatment effect exists as the effect of experimental versus control treatment is diluted by the biomarker-based treatment selection. For this reason, a large sample size is needed to detect at least a small overall difference in outcomes between the two biomarker-strategy arms.
- Should be used only if you want to evaluate a complex biomarker-guided strategy with a variety of treatment options or biomarker categories.
Key references
- Gosho, M.; Nagashima, K.; Sato, Y. Study designs and statistical analyses for biomarker research. Sensors 2012, 12, 8966–8986. [Google Scholar] [CrossRef] [PubMed]
- Beckman, R.A.; Clark, J.; Chen, C. Integrating predictive biomarkers and classifiers into oncology clinical development programmes. Nat. Rev. Drug Discov. 2011, 10, 735–748. [Google Scholar] [CrossRef] [PubMed]
- Young, K.Y.; Laird, A.; Zhou, X.H. The efficiency of clinical trial designs for predictive biomarker validation. Clin. Trials 2010, 7, 557–566. [Google Scholar] [CrossRef] [PubMed]
- Hoering, A.; Leblanc, M.; Crowley, J.J. Randomized phase III clinical trial designs for targeted agents. Clin. Cancer Res. 2008, 14, 4358–4367. [Google Scholar] [CrossRef] [PubMed]
- Kelloff, G.J.; Sigman, C.C. Cancer biomarkers: Selecting the right drug for the right patient. Nat. Rev. Drug Discov. 2012, 11, 201–214. [Google Scholar] [CrossRef] [PubMed]
- Tajik, P.; Zwinderman, A.H.; Mol, B.W.; Bossuyt, P.M. Trial designs for personalizing cancer care: A systematic review and classification. Clin. Cancer Res. 2013, 19, 4578–4588. [Google Scholar] [CrossRef] [PubMed]
- Galanis, E.; Wu, W.; Sarkaria, J.; Chang, S.M.; Colman, H.; Sargent, D.; Reardon, D.A. Incorporation of biomarker assessment in novel clinical trial designs: Personalizing brain tumor treatments. Curr. Oncol. Rep. 2011, 13, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Freidlin, B.; McShane, L.M.; Korn, E.L. Randomized clinical trials with biomarkers: Design issues. J. Natl. Cancer Inst. 2010, 102, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.R.; Galanis, E. Incorporation of prognostic and predictive factors into glioma clinical trials. Curr. Oncol. Rep. 2013, 15, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Di Maio, M.; Gallo, C.; De Maio, E.; Morabito, A.; Piccirillo, M.C.; Gridelli, C.; Perrone, F. Methodological aspects of lung cancer clinical trials in the era of targeted agents. Lung Cancer 2010, 67, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Scheibler, F.; Zumbé, P.; Janssen, I.; Viebahn, M.; Schröer-Günther, M.; Grosselfinger, R.; Hausner, E.; Sauerland, S.; Lange, S. Randomized controlled trials on pet: A systematic review of topics, design, and quality. J. Nucl. Med. 2012, 53, 1016–1025. [Google Scholar] [CrossRef] [PubMed]
- Bria, E.; Di Maio, M.; Carlini, P.; Cuppone, F.; Giannarelli, D.; Cognetti, F.; Milella, M. Targeting targeted agents: Open issues for clinical trial design. J. Exp. Clin. Cancer Res. 2009, 28, 66. [Google Scholar] [CrossRef] [PubMed]
- Renfro, L.A.; Mallick, H.; An, M.-W.; Sargent, D.J.; Mandrekar, S.J. Clinical trial designs incorporating predictive biomarkers. Cancer Treat. Rev. 2016, 43, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Ondra, T.; Dmitrienko, A.; Friede, T.; Graf, A.; Miller, F.; Stallard, N.; Posch, M. Methods for identification and confirmation of targeted subgroups in clinical trials: A systematic review. J. Biopharm. Stat. 2016, 26, 99–119. [Google Scholar] [CrossRef] [PubMed]
- Eng, K.H. Randomized reverse marker strategy design for prospective biomarker validation. Stat. Med. 2014, 33, 3089–3099. [Google Scholar] [CrossRef] [PubMed]
- Baker, S.G. Biomarker evaluation in randomized trials: Addressing different research questions. Stat. Med. 2014, 33, 4139–4140. [Google Scholar] [CrossRef] [PubMed]
- Cree, I.A.; Kurbacher, C.M.; Lamont, A.; Hindley, A.C.; Love, S. A prospective randomized controlled trial of tumour chemosensitivity assay directed chemotherapy versus physician’s choice in patients with recurrent platinum-resistant ovarian cancer. Anti-Cancer Drugs 2007, 18, 1093. [Google Scholar] [CrossRef] [PubMed]
- Cobo, M.; Isla, D.; Massuti, B.; Montes, A.; Sanchez, J.M.; Provencio, M.; Viñolas, N.; Paz-Ares, L.; Lopez-Vivanco, G.; Muñoz, M.A.; et al. Customizing cisplatin based on quantitative excision repair cross-complementing 1 mRNA expression: A phase III trial in non-small-cell lung cancer. J. Clin. Oncol. 2007, 25, 2747. [Google Scholar] [CrossRef] [PubMed]
- Lijmer, J.G.; Bossuyt, P.M.M. Various randomized designs can be used to evaluate medical tests. J. Clin. Epidemiol. 2009, 62, 364. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.-J. Biomarker as a classifier in pharmacogenomics clinical trials: A tribute to 30th anniversary of PSI. Pharm. Stat. 2007, 6, 283–296. [Google Scholar] [CrossRef] [PubMed]
- Cho, D.; McDermott, D.; Atkins, M. Designing clinical trials for kidney cancer based on newly developed prognostic and predictive tools. Curr. Urol. Rep. 2006, 7, 8–15. [Google Scholar] [CrossRef] [PubMed]